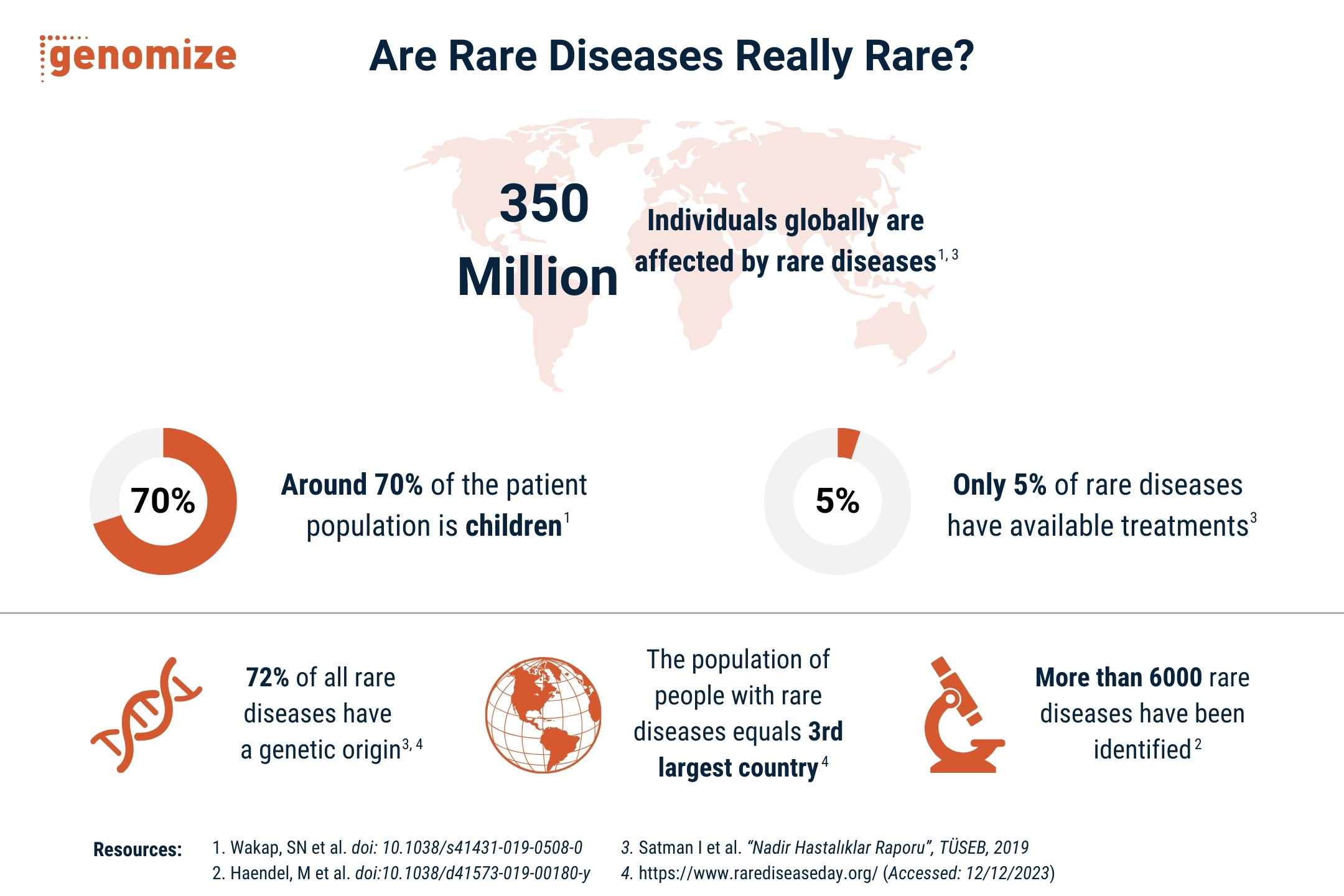
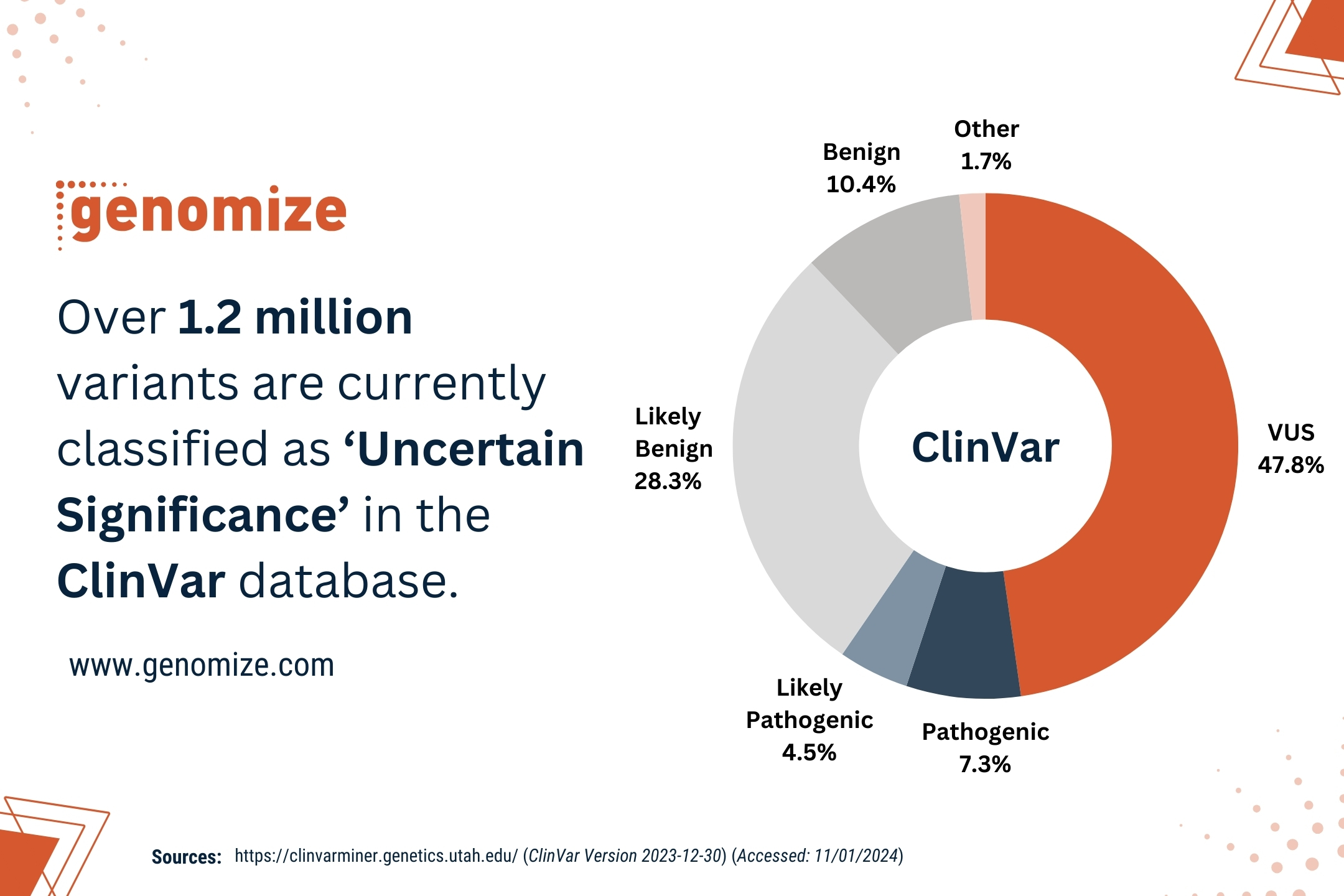
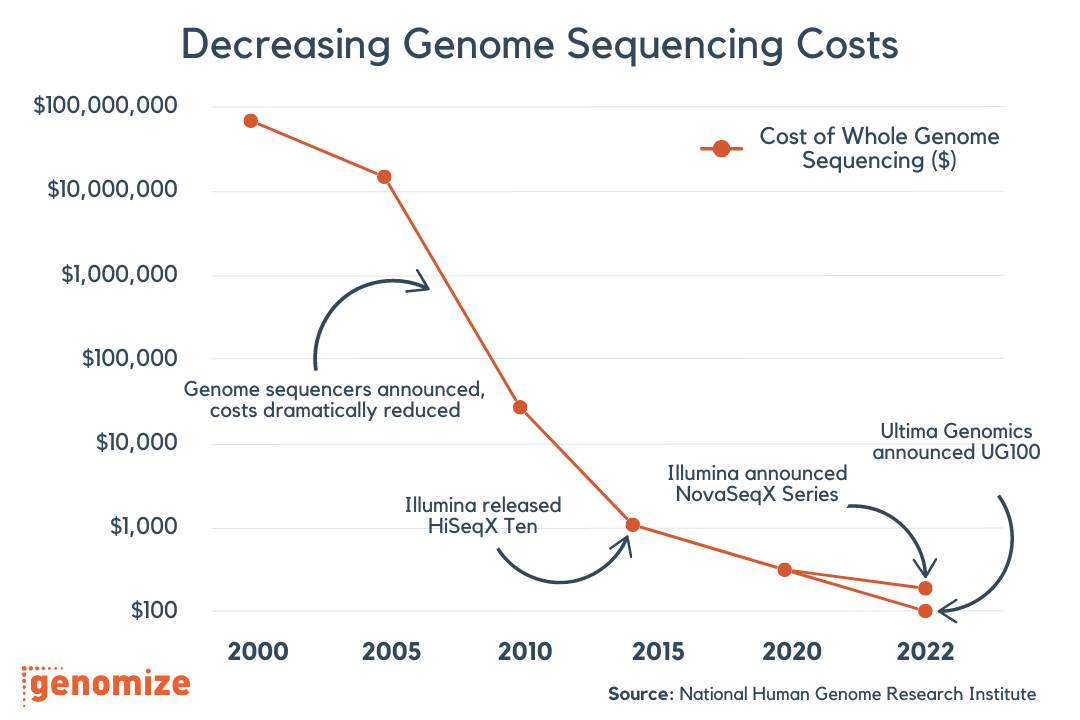
Diagnostic yields for short-read genome sequencing in rare disease cohorts typically range from 25% to 50%, depending on the patient population and inclusion criteria [1, 2, 3]. This means that even after whole-genome sequencing, more than half of individuals with a suspected genetic condition receive no molecular diagnosis. Some of these unresolved cases will involve non-genetic causes, polygenic contributions, or mechanisms that current knowledge simply cannot explain [4]. However, a meaningful proportion of these patients do carry detectable monogenic variants [1, 4].
Short-read sequencing, for all its maturity and throughput, has well-documented blind spots: structural variants with breakpoints in repetitive regions, tandem repeat expansions, phasing of compound heterozygous variants, and epigenetic modifications [5]. These are variant classes that disproportionately affect known disease genes, and they are precisely the categories where long-read sequencing has begun to demonstrate measurable diagnostic improvements.
What Short-Read Sequencing Can’t See
Short-read sequencing platforms generate reads of 50-300 base pairs [6, 7]. For single nucleotide variants (SNVs) and small insertions or deletions (indels), this is usually sufficient. However, several clinically important variant classes such as structural variants (SVs) and tandem repeat expansions fall outside of what short reads can reliably detect.
Table 1. Short-read versus long-read sequencing performance across six clinically relevant variant classes.
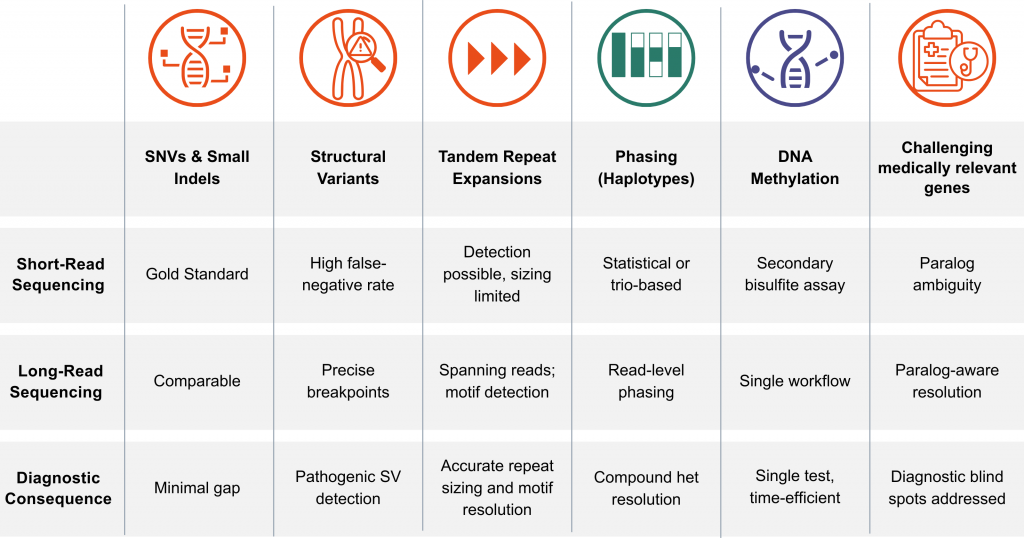
These gaps have direct clinical consequences and long-read sequencing addresses each of them directly. For example, in one critically ill infant, long-read sequencing phased compound heterozygous PLA2G6 variants in trans from a singleton sample, confirming an autosomal recessive diagnosis without parental sequencing [8].
In another case, a DNA methylation signature established a diagnosis of CHARGE syndrome (coloboma, heart defects, atresia of the choanae, retardation of growth, genital and ear abnormalities). This led the clinicians to perform a targeted CHD7 analysis, which confirmed a frameshift variant within 48 hours [8].
In a prospective cohort using long-read sequencing as a first-line test, a pathogenic FMR1 expansion was sized at 654 CGG repeats. The analysis also revealed loss of AGG interrupting motifs and confirmed concurrent promoter hypermethylation, all from a single sequencing run [9].
Long-Read WGS: What the Clinical Evidence Shows
The clinical evidence supports the use of long-read sequencing as a second-line rescue tool, a first-line diagnostic test, and a time-critical intervention.
The “Second-Line” Rescue Yield
When long-read sequencing has been applied to patients who had already received negative or inconclusive results from the short-read sequencing based tests, it has consistently increased the diagnostic output.
Across a synthesis of more than 50 studies, the added yield ranged from 7% to 17% following negative short-read WGS [9]. Individual cohort studies reflect this pattern: 16.7% of short-read-negative neurodevelopmental disorder probands received new findings with long-read sequencing, including variants uniquely resolved by the technology and cases solved through reanalysis against updated gene-disease knowledge [11]. Among patients with inconclusive exome results, the added yield was 10%, including a methylation-based diagnosis at the spinal muscular atrophy locus [12]. The gains were concentrated in structural variants, tandem repeat expansions, and phasing-dependent diagnoses, precisely the variant classes described above as outside short-read resolution.
The Move to First-Line Diagnostics
Studies evaluating long-read sequencing as a primary investigation tool were also promising. Ek et al. (2026) performed long-read sequencing and short-read sequencing in parallel for 100 prospective pediatric cases [9]. While both achieved a 29% baseline yield, long-read sequencing provided added clinical value in 14% of cases through:
- Direct phasing of compound heterozygous variants.
- Resolution of complex structural rearrangements.
- Methylation-based confirmation of imprinting disorders.
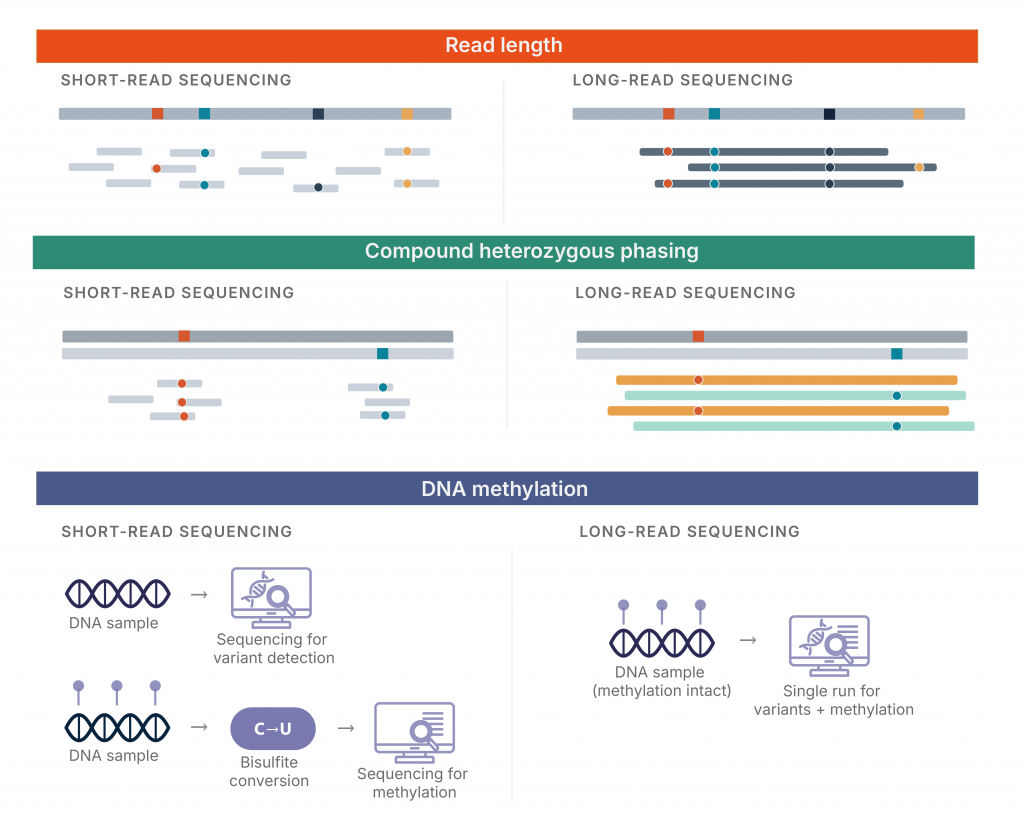
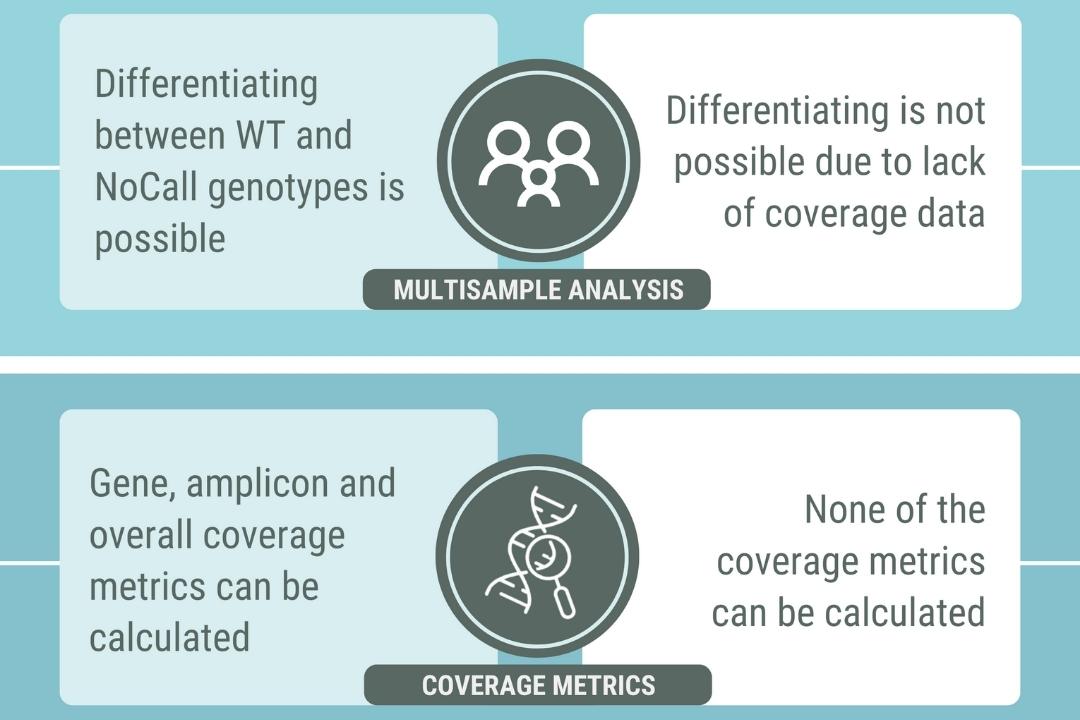
Figure 1. Three technical differences between short-read and long-read sequencing.
The authors estimated that 24% of parental follow-up tests could have been omitted had long-read sequencing been the sole test, representing a significant reduction in clinical workflow complexity.
Ultra-Rapid Diagnostics in Critical Care
In acute settings the turnaround matters as much as the yield. Smits et al. (2026) applied ultra-rapid long-read sequencing to 26 critically ill patients, predominantly neonates and children (median age 2 months) [8].
- Diagnostic Rate: 42% (11/26 cases).
- Turnaround Time: Average 5.3 days for long-read sequencing vs. 18.4 days for standard care.
- Clinical Impact: In 7 of the 11 solved cases, the long-read sequencing results led to an immediate change in management, such as medication switches or redirection of care.
Managing Complexity: The Role of Genomize SEQ Platform
As long-read sequencing addresses the technical blind spots described above, it introduces a new challenge: data volume and interpretive complexity. Long-read sequencing data differs from short-read sequencing output in ways that standard tertiary analysis pipelines were not designed for: thousands of structural variant calls per genome, size-resolved tandem repeat expansions requiring locus-specific interpretation, methylation calls integrated with sequence, and long-range phased haplotypes. Handling this at clinical scale requires interpretation platforms built for the data type. Genomize SEQ platform was built for exactly this.
Genomize SEQ Platform was developed as a clinical genomic analysis and interpretation platform; it provides the tertiary analysis layer to handle long-read whole-genome data at scale. The platform’s long-read analysis pipelines use dedicated tools for structural variants, tandem repeats, and paralog-aware resolution of challenging medically relevant genes [13] such as SMN1/SMN2, HBA1/HBA2, PMS2, and CYP21A2. Then, it automates the classification of variants according to ACMG/ClinGen guidelines, while integrating native phasing data to resolve compound heterozygosity in real time. By simplifying the path from complex genomic signals to a final molecular report, SEQ Platform enables laboratories to leverage the full diagnostic power of long-read technology without increasing their interpretive burden.
Conclusion
Evidence from recent prospective cohorts suggests that the current diagnostic gap reflects specific technical boundaries in sequencing hardware that persist even alongside the established utility of short-read platforms. While short-read sequencing has provided the foundation for modern genetics, its inherent blind spots in repetitive and structural regions present a clear ceiling in a first-line diagnostic setting (Figure 1). Long-read sequencing represents a necessary evolution beyond second-line testing, providing a more comprehensive, phased, and epigenetically aware view of the human genome. The maturation of interpretation platforms like Genomize SEQ supports this transition, positioning long-read sequencing as a practical and increasingly viable framework for clinical genomics.
References
1. Eisfeldt J, Ek M, Nordenskjöld M, Lindstrand A. Toward clinical long-read genome sequencing for rare diseases. Nat Genet. 2025;57(6):1334–43. https://doi.org/10.1038/s41588-025-02160-y
2. The 100,000 Genomes Project Pilot Investigators. 100,000 Genomes Pilot on Rare-Disease Diagnosis in Health Care — Preliminary Report. N Engl J Med. 2021;385(19):1868–80. https://doi.org/10.1056/NEJMoa2035790
3. Stranneheim H, Lagerstedt-Robinson K, Magnusson M, Kvarnung M, Nilsson D, Lesko N, et al. Integration of whole genome sequencing into a healthcare setting: high diagnostic rates across multiple clinical entities in 3219 rare disease patients. Genome Med. 2021;13(1):40. https://doi.org/10.1186/s13073-021-00855-5
4. Marwaha S, Knowles JW, Ashley EA. A guide for the diagnosis of rare and undiagnosed disease: beyond the exome. Genome Med. 2022;14(1):23. https://doi.org/10.1186/s13073-022-01026-w
5. Mantere T, Kersten S, Hoischen A. Long-read sequencing emerging in medical genetics. Front Genet. 2019;10:426. https://doi.org/10.3389/fgene.2019.00426
6. Illumina. Sequencing platforms [Internet]. [cited 2026 Apr 28]. Available from: https://emea.illumina.com/systems/sequencing-platforms.html
7. Illumina. NovaSeq 6000 sequencing system [Internet]. [cited 2026 Apr 28]. Available from: https://emea.illumina.com/systems/sequencing-platforms/novaseq.html
8. Smits DJ, Ferraro F, Drost M, et al. Nanopore long-read sequencing for the critically ill facilitates ultrarapid diagnostics and urgent clinical decision making. Eur J Hum Genet. 2026;34:108–18. https://doi.org/10.1038/s41431-025-01959-x
9. Ek M, Kvarnung M, ten Berk de Boer E, Lindstrand A. Long-read genome sequencing enhances diagnostics of pediatric neurological disorders. Genome Med. 2026;18(1):12. https://doi.org/10.1186/s13073-025-01596-5
10. Del Gobbo GF, Boycott KM. The additional diagnostic yield of long-read sequencing in undiagnosed rare diseases. Genome Res. 2025;35(4):559–71. https://doi.org/10.1101/gr.279970.124
11. Hiatt SM, Lawlor JMJ, Handley LH, Latner DR, Bonnstetter ZT, Finnila CR, et al. Long-read genome sequencing and variant reanalysis increase diagnostic yield in neurodevelopmental disorders. Genome Res. 2024;34(10):1747–62. https://doi.org/10.1101/gr.279227.124
12. Sinha S, Rabea F, Ramaswamy S, Chekroun I, El Naofal M, Jain R, et al. Long read sequencing enhances pathogenic and novel variation discovery in patients with rare diseases. Nat Commun. 2025;16:2500. https://doi.org/10.1038/s41467-025-57695-9
13. Wagner J, Olson ND, Harris L, McDaniel J, Cheng H, Fungtammasan A, et al. Curated variation benchmarks for challenging medically relevant autosomal genes. Nat Biotechnol. 2022;40:672–80. https://doi.org/10.1038/s41587-021-01158-1